Table of Contents
Retinotopy Tutorial (Scripting)
This page show how the analysis steps done in the Retinotopy tutorial can be scripted without any GUI interaction. If you don't know how to run a retinotopy analysis with the GUI then you should go through the Retinotopy tutorial.
Overview
- Download the tutorial files
- Make the directory structure for mrLoadRet and move the files into the correct locations
- Run mrInit to initialize the session
- Run the Motion Compensation
- Average wedge and ring scans together
- Run the correlation analysis
- Getting data
1. Download files
You can download the tutorial files from:
Note that this is a rather big file, approximately 300MB.
The files are provided as a tar/zip file. In Mac OS X you should be able to just click to open the files in the Finder. Otherwise, you can go to a terminal and do:
gunzip retinotopyTutorial.tar.gz tar xvf reinotopyTutorial.tar
2. mrLoadRet directory structure
Now that you have the raw image files, you need to make the directory structure for mrLoadRet and copy your files into the correct directories. This is the same steps that you preformed in the retinotopy tutorial. (Note that this does not have to be done in Matlab, it can be done from the command line or from the Finder or from a shell script that is customized for the file types at your site).
- Start by changing directories into the downloaded tutorial data directory
cd retinotopyTutorial
- Now make a directory for your anatomy files
mkdir Anatomy movefile inplane.* Anatomy movefile volume.* Anatomy movefile right_occipital.* Anatomy
- Now make the directory to hold your raw time series (i.e. the epi scans with your data), and move the scan files into that directory
mkdir Raw mkdir Raw/TSeries movefile Scan* Raw/TSeries
You should now have two top level directories (Anatomy and Raw) and all the files should have been moved into one of the sub-directories:
>> ls Anatomy/ Raw/
3. mrInit
Next, we will run mrInit without bringing up the GUI. We first get default parameters. Note that if you set defaultParams=0, then it will bring up the GUI allowing you to set parameters with the gui, but save the settings into the parameter structures rather than running mrInit.
[sessionParams groupParams] = mrInit([],[],'justGetParams=1','defaultParams=1')
Then, let's set the session information
sessionParams.description = 'Retinotopy Tutorial'; sessionParams.subject = 'Slothrop'; sessionParams.operator = 'Laszlo Jamf';
Then let's set the group parameters:
groupParams.junkFrames(:) = 8;
groupParams.nFrames(:) = 160;
groupParams.description{1} = 'CW';
groupParams.description{2} = 'CCW';
groupParams.description{3} = 'Expanding';
groupParams.description{4} = 'Contracting';
groupParams.description{5} = 'CW';
groupParams.description{6} = 'CCW';
groupParams.description{7} = 'Expanding';
groupParams.description{8} = 'Contracting';
groupParams.description{9} = 'CW';
groupParams.description{10} = 'CCW';
Now, let's run mrInit
mrInit(sessionParams,groupParams,'makeReadme=0');
4. motion compensation
Motion compensation can be fully automated, these files have been fully motion compensated already, so we will just how to set up the scripting, but not actually do it.
First get a view.
v = newView;
Now get default params
[v params] = motionComp(v,[],'justGetParams=1','defaultParams=1');
You can change any of the params that you like
>> params
params =
groupName: 'Raw'
baseScan: 1
baseFrame: 'first'
sliceTimeCorrection: 1
sliceTimeString: 'middle of TR'
robust: 0
correctIntensityContrast: 0
crop: []
niters: 3
motionCompGroupName: 'MotionComp'
interpMethod: 'linear'
targetScans: [1 2 3 4 5 6 7 8 9 10]
tseriesfiles: {1x10 cell}
descriptions: {1x10 cell}
tSmooth: 0
Once you have the params the way you want them to be, then you can run the motionComp with (but don't do this, since it will take a long time and it has already been done):
v = motionComp(v,params);
5. Average time series
Start by getting a view:
v = newView;
First we get parameters for averaging together all the wedge scans. Start by getting the scan numbers for the CW and CCW wedge scans (We already know the scan numbers because we explicitly set which scan was which above, but it is always a good idea not to assume anything about scan numbers of group numbers).
CWScanNums = viewGet(v,'scanNumFromDescription','CW','Raw','exact'); CCWScanNums = viewGet(v,'scanNumFromDescription','CCW','Raw','exact'); scanList = [CWScanNums CCWScanNums];
The scanList should look like the following:
>> scanList
scanList =
1 5 9 2 6 10
Now get the parameters for the averaging:
[v params] = averageTSeries(v,[],'justGetParams=1','defaultParams=1','scanList',scanList);
Now, we make sure to set the shift and reverse list approriately:
params.shiftList(:) = 2; params.reverseList(1:length(CWScanNums)) = 1;
Then run the averaging:
averageTSeries(v,params);
Now do the same thing for the ring stimuli:
expandingScanNums = viewGet(v,'scanNumFromDescription','Expanding','Raw','exact'); contractingScanNums = viewGet(v,'scanNumFromDescription','Contracting','Raw','exact'); scanList = [expandingScanNums contractingScanNums]; [v params] = averageTSeries(v,[],'justGetParams=1','defaultParams=1','scanList',scanList); params.shiftList(:) = 2; params.reverseList(1:length(expandingScanNums)) = 1; averageTSeries(v,params);
We can get back structures that will have fields containing all the information we need to confirm that we did everything properly.
gInfo = groupInfo;
gAverageInfo = groupInfo('Averages');
sInfo = scanInfo(1,'Averages');
This structures will then have:
>> gInfo(end)
ans =
groupName: 'Averages'
numScans: 2
dirSize: 143024
>> gAverageInfo(end)
ans =
description: 'Average from Raw of scans: 3 7 4 8'
scanVoxelSize: [3.00104141235352 3.0054624080658 2.99838328361511]
tr: 1.5
totalFrames: 160
filename: 'tseries-081101-233216.img'
originalFilename: {'Scan03_EXP.img' 'Scan04_CON.img' 'Scan07_EXP.img' 'Scan08_CON.img'}
originalGroupname: {'Raw' 'Raw' 'Raw' 'Raw'}
stimFilename: {}
scanDims: [64 64 27]
totalJunkedFrames: [0 0 0 0]
junkFrames: 0
>> sInfo
sInfo =
description: 'Average from Raw of scans: 1 5 9 2 6 10'
Filename: 'tseries-081101-233027.img'
GroupName: 'Averages'
Original1: 'Raw: Scan01_CW.img'
Original2: 'Raw: Scan05_CW.img'
Original3: 'Raw: Scan09_CW.img'
Original4: 'Raw: Scan02_CCW.img'
Original5: 'Raw: Scan06_CCW.img'
Original6: 'Raw: Scan10_CCW.img'
voxelSize: [3.00104141235352 3.0054624080658 2.99838328361511]
dims: [64 64 27]
TR: ''
framePeriod: 1.5
numVolumes: 160
junkFrames: 0
totalJunkedFrames: '0 0 0 0 0 0'
qform: [4x4 double]
qformCode: 1
sform: [4x4 double]
sformCode: 0
vol2mag: ''
vol2tal: ''
scanList: '1 5 9 2 6 10'
shiftList: '2 2 2 2 2 2'
reverseList: '1 1 1 0 0 0'
baseScan: '1'
interpMethod: 'nearest'
paramInfo: {1x27 cell}
Which should be info that you can check in a script.
Finally, let's change the description of the averaged scans. Note that the two averaged scans should be the last two scans in the Average group (i.e. nScans-1 and nScans):
nScans = viewGet(v,'nScans','Averages');
scanParams = viewGet(v,'scanParams',nScans-1,'Averages');
scanParams.description = sprintf('Wedges: %s',scanParams.description);
v = viewSet(v,'scanParams',scanParams,nScans-1,'Averages');
scanParams = viewGet(v,'scanParams',nScans,'Averages');
scanParams.description = sprintf('Rings: %s',scanParams.description);
v = viewSet(v,'scanParams',scanParams,nScans,'Averages');
Finally, save the session to make sure that the changes above stick
saveSession
6. Correlation analysis
You can script the correlation analysis by first changing to the Averages group and then getting parameters:
v = viewSet(v,'curGroup','Averages'); [v params] = corAnal(v,[],'justGetParams=1','defaultParams=1');
Then set the parameters appropriately:
params.recompute(:) = 1; params.ncycles(:) = 10;
Now run the corAnal
corAnal(v,params);
Finally, if you are done using the view variable, remember to delete it.
deleteView(v);
There. We are done. You have now done the same analysis as you did using the GUI in the retinotopy tutorial. You can now run mrLoadRet and view the results in the same way as you did with the tutorial. After you open up the mrLoadRet viewer, make sure you are switched to the Averages Group and use File/Analysis/Load to load the corAnal/corAnal.mat.
7. Getting data
Getting overlay values
Instead of running the GUI you may actually just want to grab the coherence, amplitude and phase data from the analysis for your own analysis program. You can do that by getting the information from a view variable. You can either use the view variable you were using above, or start a new one:
v = newView;
Now switch to the Averages group and find the wedges scan:
v = viewSet(v,'curGroup','Averages'); wedgesScanNum = viewGet(v,'scanNumFromDescription','Wedges'); v = viewSet(v,'curScan',wedgesScanNum);
Load the computed corAnal analysis in:
v = loadAnalysis(v,'corAnal/corAnal.mat');
Let's get the phase overlay:
overlayNames = viewGet(v,'overlayNames');
phOverlayNum = find(strcmp('ph',overlayNames));
v = viewSet(v,'curOverlay',phOverlayNum);
phOverlay = viewGet(v,'overlayData',wedgesScanNum);
phOverlay now is a 64x64x27 matrix that contains the phase value for every voxel in the volume.
Make sure to remember to delete your view when done:
deleteView(v);
Getting overlay image
You can also get an overlay image like the one displayed in the viewer, without having to interact with the viewer GUI. To do this, start up MLR:
mrLoadRet([])
The empty brackets makes it forget previous settings. Now, change the group and scan to the wedges scan like before, but instead of using a view you created, use the MLR view that is returned by getMLRView:
viewSet(getMLRView,'curGroup','Averages'); wedgesScanNum = viewGet(getMLRView,'scanNumFromDescription','Wedges'); viewSet(getMLRView,'curScan',wedgesScanNum);
Load the analysis
loadAnalysis(getMLRView,'corAnal/corAnal.mat');
Load the right_occipital anatomy:
loadAnat(getMLRView,'right_occipital.hdr');
Set the overlay to ph
overlayNames = viewGet(getMLRView,'overlayNames');
phOverlayNum = find(strcmp('ph',overlayNames));
viewSet(getMLRView,'curOverlay',phOverlayNum);
Now refresh the MLR display and grab the image
img = refreshMLRDisplay(viewGet(getMLRView,'viewNum'));
ok, now display
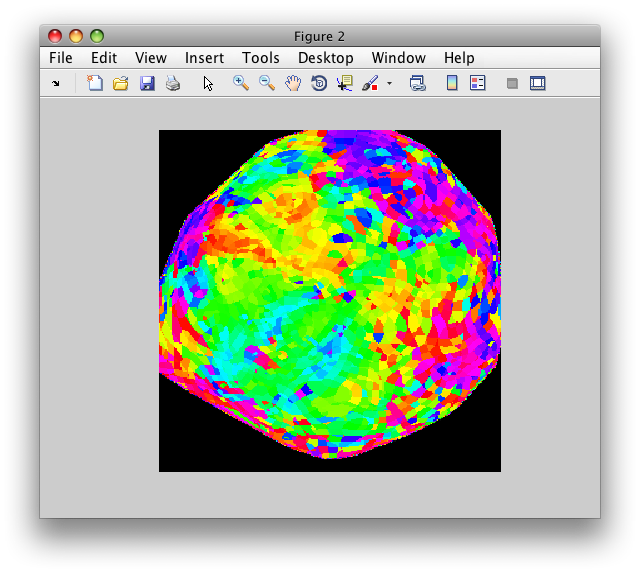
figure; image(img); axis square; axis off;
And you should get this:
To close the viewer:
mrQuit(0);