Table of Contents
Function Reference
This is not a complete list of functions. This contains references to some of the most useful functions in the mrLoadRet distribution.
Core mrLoadRet functions
mrInit
usage: [sessionParams groupParams] = mrInit(<sessionParams>,<groupParams>,<justGetParams=1>,<defaultParams=1>)
purpose:This is a replacement for mrInitRet, which can be localized easily for processing data collected at other sites.
| argument | value |
|---|---|
| <sessionParams> | Optional argument used when scripting. |
| <groupParams> | Optional argument used when scripting. |
| <justGetParams=1> | Set this to get parameters w/out running. |
| <defaultParams=1> | Set this to use default parameters. |
The return arguments are
| return argument | value |
|---|---|
| sessionParams | Session parameters. |
| groupParams | Group parameters. |
cd to the folder containing the session you want to analyze. Inside this folder you should have made the following subfolders:
- Anatomy Contains your 2D inplane anatomy file (or any other anatomy you want to display your data on).
- Raw A directory for the raw scan files from the scanner. Underneath this folder, you will need to create a subfolder TSeries (i.e. Raw/TSeries) which contains all of your scan files in nifti file. Note that the default is to use the .hdr/.img nifti pair, but you can override this by setting the niftiFileExtension parameter from the mrLoadRet menu item Edit→Preferences (or from the command line using: mrEditPrefs) to .nii.
- Etc A directory containing miscellaneous files. This is where you should put “stimfiles” from mgl if you are using mgl. Or eye tracker files. Or anything else associated with the scan.
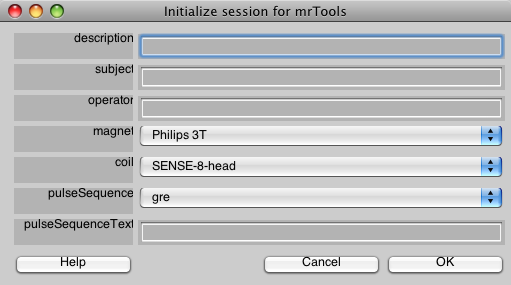
Call mrInit at the matlab prompt and fill in the information in the GUI windows: a brief description, subject, operator, and information about scanner and sequences. Make sure that the directory structure inside the current folder for your session is set up correctly before (usual default is ssYYYYMMDD/Raw/TSeries, ssYYYYMMDD/Anatomy, where subject initials and date in year-month-day is a common convention).
By default mrInit also calls mrReadme which produces a text file that contains basic information about the scanning session. This is useful for finding a particular scanning session some time after you have collected the data, using e.g. at the OSX command line with grep
Say, all your data sets are contained in a folder called /data/projects, then you can look for the descriptions of all the data sets in that directory by:
$ grep --recursive --include Readme.txt 'Description:' /data/projects
Site specific changes
By setting preferences in your startup file you can adjust the options in the GUI dialog boxes to suit your needs. Specifically, you might want to change the preferences magnet, coil, and pulseSequence.
Here are a couple of example additions to the startup.m file for Site:Nottingham
mrSetPref('site','Nottingham');
...
mrSetPref('magnet',{'Philips 3T', 'Philips 7T', 'other'} );
mrSetPref('coil',{'SENSE-8-head','FlexS', 'SENSE-16-head', 'other'} );
mrSetPref('pulseSequence',{'gre', 'se', 'other'} );
Localized changes in pull-down menus
viewGet/viewSet
Whenever you need a piece of information, always get it with a viewGet. Do not try to dereference global structures like MLR or the view variable returned by newView. This way if something is changed about the way something is stored in the structures your code will not get broken.
Likewise if you have to add something (like an analysis or overlay etc). Always use an appropriate viewSet call so that it side-effects the global variable correctly and does appropriate checks.
viewGet
usage: val = viewGet(v,param,varargin)
purpose: viewGet allows you to get lots of information about your scan.
| argument | value |
|---|---|
| v | The view from which you want to get information. |
| parameterName | The name of the parameter you want information. |
| varargin | There are variable numbers of arguments passed in depending on what parameter you are getting. |
viewGet is the primary way to get information about the scan. You should always use viewGet rather than going searching around in the global variables for information you need–that way if the implementation changes in the future you won't be stuck with dereferencing variables that may have changed.
You can get many, many pieces of information. To see a full listing, type viewGet without any arguments:
viewGet
To get specific help about a particular parameters:
viewGet([],'scanNum','help')
viewSet
usage: v = viewSet(v,param,val,varargin)
purpose: viewSet allows you to get set information about the scan.
| argument | value |
|---|---|
| v | The view for which you want to set the parameter value. |
| parameterName | The name of the parameter you want to set. |
| value | The value to set that parameter to. |
Like viewGet, you should always use this function rather than trying to set global variables yourself. This will always properly side-effect the global variables. To get a list of things you can viewSet, type viewSet without any arguments:
viewSet
To get specific help about a particular parameters:
viewSet([],'scanNum','help')
Always remember to accept the returned view variable from viewSet (except if you are using getMLRView):
v = viewSet(v,'parameterName',value);
refreshMLRDisplay
usage: [img] = refreshMLRDisplay(viewNum);
purpose: This is the function that displays the image that you see in the viewer.
| argument | value |
|---|---|
| viewNum | The viewNum of the view you wish to refresh. |
refreshMLRDisplay takes care of computing the overlay and the location of the ROIs and displays that. This is called by the GUI code and you do not normally need to call it yourself. The one exception is if you are interested in getting back a matrix in which the image found in the viewer is stored. You can do this by running:
img = refreshMLRDisplay(viewGet(getMLRView,'viewNum'));
newView
usage: v = newView;
purpose: Initializes a new view variable.
Use this function to get a “view” which allows you to access all the information about your session. Many of the functions require a view as an input argument. When you are done using a view, you should use deleteView or mrQuit to remove the view. It is recommended not to use the variable name “view” for your view since that is a Matlab command.
deleteView
usage: deleteView(v);
purpose: Deletes a view variable
| argument | value |
|---|---|
| v | view variable returned by newView |
When you are finished with the view, delete it using this function
mrQuit
usage: mrQuit(<saveMrLastView>)
purpose: Quits all open views, including the MLR viewer (if one is opened).
| argument | value |
|---|---|
| saveMrLastView | Set to 0 if you don't want to save the MLR viewer settings in mrLastView (defaults to 1) |
Function used to quit all open views including the MLR viewer. This is called from the quit menu item, but if you call from the command line directly without any arguments it will close all open views and the viewer. Note that if you have multiple viewers open on the data set, it will use the last view opened to save settings in mrLastView:
mrQuit
Set saveMrLastView to 0 if you do not want to save a mrLastView. i.e.
mrQuit(0);
Time Series
loadTSeries
usage: tSeries = loadTSeries(v,[scan],[slice],[frame],[x],[y],[precision])
purpose: Loads the time series for the scan
| argument | value |
|---|---|
| v | The view variable |
| scan | Number specfying which scan to load |
| slice | either a number specifying which slice to load or 'all' which loads all of the slices. Default: 'all' |
| frame | optional number specifying which temporal frame to load. Ignored if slice is specified. Default: [] (load all frames). |
| x | Which x points to load |
| y | which y points to load |
| precision | precision of data returned, default is 'double' can be 'single' |
Loads the tSeries corresponding to the specified scan and slice. The tseries files must be in <homedir>/<view>/<group>/TSeries, and must be nifti (or analyze) format (4D file containing all of the slices and temporal frames from a given scan).
| return argument | value |
|---|---|
| tSeries | If a single slice is specified, tseries is returned as a 2D matrix (nFrames x nVoxels). If slice is 'all' then the tseries is returned as a 4D array [x y z t]. If frame is specified, then the tseries is a single volume returned as a 3D array [x y t] |
loadROITSeries
usage: rois = loadROITSeries(v,<roiname>,<scanList>,<groupNum>)
purpose: Loads ROI variables that have the tSeries loaded
| argument | value |
|---|---|
| v | The view variable |
| roiname | The name of the roi to load. May be a cell array of roinames. If roiname is a roi struct instead of a name then it will use that roi instead of loading the roi from disk. Also, if roiname is a number or cell array of numbers then it will use the corresponding ROI from the view. If not passed in, will bring up a selection dialog |
| scanList | List of scans to load tSeries for |
| groupNum | Group num to load data from |
The returned variable (or cell array if multiple ROIs are loaded) will be an ROI structure with a tSeries field added on. It will also have a scanCoords field which specifies the coordinates from the scan that were used in getting the tSeries.
For example, to load the first scan from the first group:
v = newView rois = loadROITSeries(v,'l_mt',1,1);
If you want to load the coordinates w/out loading the tSeries, you can do:
v = newView rois = loadROITSeries(v,'l_mt',1,1,'loadType=none');
Loading coordinates
Load ROI Coordinates
usage: scanCoords = getROICoordinates(view,roiNum,<scanNum>,<groupNum>)
purpose: get roi coordinates in scan coordinates
| argument | value |
|---|---|
| v | The view variable |
| roiNum | The number of the ROI to load. If roiNum is a string, it will load the ROI from the directory. If roiNum is a roi struct instead of a name then it will use that roi instead of loading the roi from disk. If not passed in, will bring up a selection dialog |
| <scanNum> | The scan whose coordinates you are trying to match. if scanNum is 0, then it will compute in the current base coordinates. |
| <groupNum> | Group num from which to choose the scan |
for example:
v = newView; coordinates = getROICoordinates(v,'rightIPS');
will return the coordinates for the ROI named 'rightIPS', and these coordinates will be consistent with the current base coordinates.
whereas:
v = newView; coordinates = getROICoordinates(v,'rightIPS',1,2);
will return the coordinates for the ROI named 'rightIPS', in the coordinates that are compatible with the first scan in the second group.
Keep ROI voxel count consistent across sessions
getROICoordinatesMatching.m
usage: scanCoords = getROICoordinatesMatching(v,roiNum,scanNum,matchScanNum,<groupNum>,<matchGroupNum>)
purpose: This gets the rois scan coordinates for the scanNum/groupNum. It does this in a special way. It first gets the scan coordinates of the roi for the matchScanNum/matchGroupNum. It then finds the corresponding coordinate for each one of these coordinates in the scanNum. This is done so that the roi for the scanNum has exactly the same voxels as the matchScanNum roi. This can be useful for classification protocols in which you want to have the same voxels from different sessions. This would usually be run on two scans that have been imported from different sessions
| argument | value |
|---|---|
| v | The view variable |
| roiNum | The number of the ROI to load. If roiNum is a string, it will load the ROI from the directory. If roiNum is a roi struct instead of a name then it will use that roi instead of loading the roi from disk. If not passed in, will bring up a selection dialog |
| <scanNum> | The first scan whose coordinates you are trying to match. |
| matchScanNum | The second scan you will be matching voxel count with |
| <groupNum> | Group num from which to choose the first scan |
| <matchGroupNum> | Group num from which to choose the second scan. If matchGroupNum is not specified, it is assumed to be the same as the groupNum. |
for example:
v = newView; Coords = getROICoordinatesMatching(v,'leftIPS',1,1,1,2);
would get the coordinates of the ROI named leftIPS, so that there are the same number of voxels in the ROI for the first scans of group 1 (perhaps scans from Monday) and the first scan from group 2 (perhaps scans from Tuesday).
Custom analyses
mrDispOverlay
usage: [view analysis] = mrDispOverlay(overlay,scanNum,groupNum/analysisStruct,<view>)
purpose: Displays an overlay in mrLoadRet. This is a quick way to get the output of your analysis into MLR.
| argument | value |
|---|---|
| overlay | A matrix the size of your scan dimensions containing the overlay you want to display |
| scanNum | The scan you want to display the overlay for |
| groupNum or AnalysisStruct | Either the groupNum to display the analysis on, or an analysis structure to insert the overlay into |
| <view> | Optional view, for when not displaying to the MLR viewer |
| varargin | A number of variable arguments are available, see below. |
Overlay is a structure that contains the overlay you want to display. It should be a 3D array with the same dimensions as the scan.
If called without a view argument it will bring up MLR.
If you pass in an analysis structure then it will add the overlay to the analysis strucutre and display that.
For example, to display a map for scan 1, group 2, where map has dimensions size(map) = [x y s]
mrDispOverlay(map,1,2)
If you want to install maps for several scans, make map into a cell array of maps of the correct dimensions and do
mrDispOverlay(map,[1 2],2);
If you want to install multiple maps for a single scan, make a cell array of maps of all the maps you want to associate with the scan and do:
mrDispOverlay(map,1,2,[],'overlayNames',{'map1name','map2name'});
If you want to install the overlay into an existing analysis, rather than a group (say erAnal), pass that analysis instead of the group:
mrDispOverlay(map,1,erAnal);
If you want to install the map into an existing view, you can pass that (or empty if you don't want to display at all):
mrDispOverlay(map,1,erAnal,[]);
If you just want to save an analysis structure without viewing anything:
v = newView; map = rand(viewGet(v,'scanDims')); scanNum = 1; groupNum = 1; mrDispOverlay(map,scanNum,groupNum,[],'overlayName=rand','saveName=randAnalysis');
If you want to return an image array that has the overlay on a specified anatomy, you could do the following:
v = newView; v = loadAnat(v,'jg_left_occipital.hdr'); map = rand(viewGet(v,'scanDims')); v = mrDispOverlay(map,1,3,v); img = refreshMLRDisplay(viewGet(v,'viewNum'));
The following is an example in which you want to set an overlay for voxels coming from some ROI.
v = newView; % load the roi (this is just to get the scan coordinates for the ROI, so we set not to load the time series) r = loadROITSeries(v,'lh_mt',1,'Concatenation','loadType=none'); % get the dimensions of the scan scanDims = viewGet(v,'scanDims',1,'Concatenation'); % Compute the linear coordinates of the roi - this makes it easier to set points in the overlay % but, you could instead write a for loop and use the x,y,z points instead... r.linearScanCoords = sub2ind(scanDims,r.scanCoords(1,:),r.scanCoords(2,:),r.scanCoords(3,:)); % create an overlay which has zeros everywhere in the scan overlay = zeros(scanDims); % and set all the voxels from the ROI to have a value of 1 overlay(r.linearScanCoords) = 1; % Then use mrDispOverlay to display mrDispOverlay(overlay,1,'Concatenation');
You can also set optional parameters:
Some parameters you can set
| parameterName | value |
|---|---|
| overlayName | name of overlay or cell array of names |
| cmap | Colormap to use, e.g. hot |
| colormapType | Can be 'setRangeToMax' |
| range | A two-array specifying the range of the overlay data, e.g. [0 1] |
| clip | A two array specifying the values to clip to, e.g. [0 1] |
| interrogator | The name of the interrogator function |
| analName | The name of the analysis to create |
| d | Installs a d structure to use for the analysis |
| params.x | A way to set arbitrary parameters, i.e. x can be any name |
| saveName | A filename for use when saving instead of displaying |
getMLRView
usage: getMLRView
purpose: To get the view variable from the current MLR Display
This function allows you to temporarily steal the view from the open mrLoadRet window. For example, if you have a mrLoadRet window open and you want to viewGet a piece of information from that window, you can do:
viewGet(getMLRView,'homeDir')
You can use it in a viewSet as well:
viewSet(getMLRView,'stimfilename','060918_stim01.mat')
Note that you should always be recalling this function to get the view. Do not set a local variable to the output of this function (i.e. v = getMLRView) and continue to use that variable. If you do, then the variable v will get out of sync with the viewer.
File I/O
cbiReadNifti
usage: [data,hdr]=cbiReadNifti(fname,subset,prec,short_nan)
purpose: Read a nifti file into matlab
| argument | value |
|---|---|
| fname | Filename of nifti file to open (needs extension) |
| subset | 4×1 cell array describing image subset to retrieve. |
| prec | Data precision |
| short_nan | How to handle NaN values for short data |
Loads all or part of a Nifti-1 file.
Default is to load entire file as a double precision array with dimensions [xSize ySize zSize tSize] where x,y,z,t are defined according to the Nifti standard (x fastest changing dimension, y second and so on - usually x corresponds to left-right, y to back-front, z to down-up, and t to early-late. NB: will also work for quasi-5-dimensional data, e.g. 3D arrays of vectors defined on a single time point (t then being the dimensions of the vectors at each point), i.e. hdr.dim(4)==1 and hdr.dim(5)>1. (0-offset index) Will not work with higher dimensional data sets (hdr.dim(4)>1 & hdr.dim(5)>1). Ignores hdr.dim(6) and higher.
hdr contains the Nifti header returned by readNiftiHeader.
Options:
- subset: 4×1 cell array describing image subset to retrieve. 1-offset (Matlab-style). Examples:
- to retrieve a single z-slice (e.g. 4): subset={[],[],4,[]}
- to retrieve a single voxel time series (e.g. [5,6,7]): subset={5,6,7,[]}
- to retrieve a single volume from a time series (e.g. second volume): subset={[],[],[],2}
- to retrieve a block of voxels from a volume: eg. subset={[4 50],[6 20],[1 10],[]}
- If data is quasi-5D, subset{4} defines the subset in dim(5)
- prec: Storage type of returned data. Legal values are:
- 'native' - data are returned as stored in file (no scaling). Only works for loading contiguous data (entire volumes).
- 'double' - data are returned in Matlab standard double precision format (default)
- 'single' - data are returned in Matlab single precision format
- short_nan: NaN handling for signed short (int16) data. If 1, will treat -32768 (smallest representable number) as NaN, reserving -32767..32767 for scaled data; otherwise treats -32786 as a real value. Default is 1.
cbiReadNiftiHeader
usage: hdr = cbiReadNiftiHeader(fname)
purpose: Read a Nifti header into Matlab
| argument | value |
|---|---|
| fname | Name of header file to open |
Loads the header from a NIFTI-1 file.
The header struct contains all stored fields plus some additional extracted fields. These are:
- .hdr_name name of header file
- .img_name name of image file
- .is_analyze 1 if valid Analyze, but not a valid NIFTI file, 0 otherwise
- .single_file flag for single file (1 for .nii, 0 for .hdr/.img)
- .endian endianness of file
- .matlab_datatype data type in Matlab string format (see fread)
- .qform44 homogeneous transform matrix extracted from qform data (rigid body - native coordinate system) ([] for Analyze)
- .sform44 homogeneous transform matrix extracted from sform data (affine - optional alignment transform) ([] for Analyze)
fname can have any legal extension (.hdr, .img, .nii). No support for compressed data – will bail out.
cbiWriteNifti
usage: [byteswritten,hdr]=cbiWriteNifti(filename,data,hdr,prec,subset,short_nan)
alternate usage: [byteswritten,hdr]=cbiWriteNifti(filename,data,[],prec);
purpose: Writes a Nifti file
| argument | value |
|---|---|
| filename | Name of file to save to |
| data | The image data to save |
| hdr | The nifti header to save, can be empty |
| prec | The precision of the data to write |
| subset | What subset of the data to write |
| short_nan | Nan handling for short values |
The header is always passed through cbiCreateNiftiHeader to ensure consistency with the data. Note in particular that if you have changed the qform44 or the sform44 field, you must call cbiSetNiftiQform and/or cbiSetNiftiSform for these changes to take effect.
- prec Precision (dataype) of output. Default (if no header is specified) is 'float32'. Use [] for default. Should be a recognized Matlab (or nifti) data type string.
- subset 4×1 cell array describing image subset to save. 1-offset (Matlab-style). Only the following options are supported:
- to save a single z-slice (e.g. 4): subset={[],[],4,[]}
- to save a single/multiple volumes of a time series (e.g. volumes 2-9): subset={[],[],[],[2 9]}
- short_nan NaN handling for signed short (int16) data. If 1, will treat save NaN's as -32768 (smallest representable number) reserving -32767..32767 for scaled data; otherwise will save NaN as 0. Default is 1 (use -32768 as NaN).
cbiSetNiftiQform
usage: hdr = cbiSetNiftiQform( hdr, mtx44 );
alternate usage: hdr=cbiNiftiSetQform( hdr, R, T );
purpose: Sets qform44 and updates quaternions, pixdim, and qoffsets accordingly. You must use this function rather then setting it by hand since this maintains consistency with the quaternion representation in the header. Also sets qform_code = 1.
| argument | value |
|---|---|
| hdr | Nifti header |
| mtx44 | 4×4 homogenous transformation matrix |
| R | 3×3 rotation matrix |
| T | A translation 3-vector |
cbiSetNiftiSform
usage: hdr = cbiSetNiftiSform( hdr, mtx44 );
alternate usage: hdr=cbiNiftiSetSform( hdr, R, T );
purpose: Sets sform44 and updates srow_x/y/z. You must use this function rather then setting it by hand since this maintains consistency with the quaternion representation in the header. Also sets sform_code = 1.
| argument | value |
|---|---|
| hdr | Nifti header |
| mtx44 | 4×4 homogenous transformation matrix |
| R | 3×3 rotation matrix |
| T | A translation 3-vector |
loadSurfOFF
usage: surf = loadSurfOFF(surffile,<loadOnlyHeader>);
purpose: Loades an off surface into matlab
| argument | value |
|---|---|
| surffile | Filename of surface in OFF binary format to load |
| loadOnlyHeader | Set to 1 to just load the header |
This code was ripped directly from Jonas Larsson's tfiReadOFF code the main difference is that this code returns a neat structure, rather than a bunch of variables
IMPORTANT NOTE Jonas' SurfRelax surfaces keep coordinates in “world” coordinates that is, they are offset by the offset in the nifti header of the volume anatomy that they were created from. To get the indexes into the volume anatomy (which is what you usually want), you have to run the function:
surf = xformSurfaceWorld2Array(surf,hdr)
where hdr is the nifti header of the volume anatomy the surface was generated from
The surf structure consists of the following variables:
- Nvtcs Number of vertices in surface
- Ntris Number of triangles in surface
- Nedges Number of edges in surface
- vtcs Nvtcs x 3 matrix of vertex coordinates
- tris Ntris x 3 matrix of triangle vertex indexes
- nParent For patches only, parent surface dimensions
- nPatch For patches only, patch surface dimensions
- patch2parent For patches only, an array that specifies the correspondence between vertices in the patch and vertices in the surface
writeOFF
usage: surf = writeOFF(surf, outName);
purpose: Saves a surface to file
| argument | value |
|---|---|
| surf | Surface structure like the one returned by loadSurfOFF |
| outName | Name of file to save as |
loadVFF
usage: [data,hdr]=loadVFF( filename, <loadOnlyHeader>)
purpose: Load a VFF file, like the ones used to store curvature in.
| argument | value |
|---|---|
| filename | Name of file to load |
| loadOnlyHeader | Set to 1 to only load header |
Loads data in .vff format. This function is borrowed from Jonas' TFI distribution. The data should be a 1xn array of curvature values (for example) for each vertex in a surface.
saveVFF
usage: saveVFF(filename,data)
purpose: Saves a file in VFF format
| argument | value |
|---|---|
| filename | Name of file to save as |
| data | A 1xn array of vertex coloring values (e.g. curvature) |
copyNiftiFile
usage: copyNiftiFile(fromFilename,toFilename,<makeLink>)
purpose: copy a nifti file. Handles copying both .hdr and .img files checks for file existence. If makeLink is set to 1, will link the files rather than copy them.
| argument | value |
|---|---|
| fromFilename | Name of nifti file to copy |
| toFilename | Name of nifti file to copy to |
| <makeLink> | Defaults to 0, set to 1 to make links rather than copy |
linkFile
usage: linkFile(fromFilename,toFilename)
purpose: links fromFilename to toFilename. Will make the link into a relative filename if possible.
| argument | value |
|---|---|
| fromFilename | Name of source file of link |
| toFilename | Name of destination file to make |
mlrFixPermissions
usage: mlrFixPermissions(dirname)
purpose: Fixes file permissions in a directory. Sets all files to be 664 and all directories to be 775.
| argument | value |
|---|---|
| dirname | Name of directory to fix permissions in |
Accessory mrLoadRet functions
importGroupScans
usage: importGroupScans
purpose: This is the same function that is available from the File/Import menu item. It allows you to import scans from another session.
mrCleanDir
usage: mrCleanDir(<force>)
purpose: Cleans up scan directories to remove unnecessary files
| argument | value |
|---|---|
| <force> | Set to 1 to clean up the directory without being asked. |
If you delete scans, the original files will still be around, even though they are unlinked in the mrSession structure. This is by design since it will keep you from deleting actual data by mistake. If you really want to delete the original files as well, run mrCleanDir. It will check for consistency with the file structure and prompt you to delete any files that are not linked. If all the files are properly matched, it will just tell you that everything maches and return:
>> mrCleanDir Group Raw matches (8:8) Group MotionComp matches (8:8)
If you don't want to be asked about whether to delete the scan files, you can set the force argument:
>> mrCleanDir(1)
makeEmptyMLRDir
usage: makeEmptyMLRDir(dirname)
purpose: Makes an empty MLR directory with a valid mrSession that can be used (for example) to import groups into.
| argument | value |
|---|---|
| dirname | Name of directory to make |
Displaying Information
groupInfo
usage: groupInfo(<groupNum>)
purpose: This function can be used to print out information about groups.
| argument | value |
|---|---|
| groupNum or groupName | Group number/name that you want info on. |
For example:
>> groupInfo('MotionComp')
1: Full motion comp of Raw scan 1: fix_center
Filename: tseries-070308-171727.img GroupName: MotionComp
Original Filename: 07+cbi_ep2d_bold_recon_swapdim+x-y.img Group: Raw
voxelSize=[3.0 3.0 3.0] TR=1.5000 Volumes=320
...
Note that this will print out information like the original filename that a group comes from.
Without any arguments, this function will print out information about all groups:
>> groupInfo
======================================== homeDir: /Volumes/BLUEBRICK/data/veins/jg060208 description: jg060208 operator: XX subject: jg magnet: Allegra 3T coil: XXX protocol: XXX ======================================== 1: Raw (7 scans) 947.9M 2: MotionComp (7 scans) 736K 3: Concatenation (5 scans) 2.5G
Note that you can specify a return argument so that you get the above information is a structure:
gInfo = groupInfo;
gInfoRaw = groupInfo('Raw');
scanInfo
usage: scanInfo(scanNum,groupNum,<displayInDialog>)
purpose: Use this function to get information about scans
| argument | value |
|---|---|
| scanNum | Number of scan you want to get information on |
| groupNum or groupName | Group number/name for the scan you want info on. |
| <displayInDialog> | Set to 1 if you want a dialog with the information. |
For example you can get a print out like:
>> scanInfo(1,'Raw') (scanInfo) Gathering scan info took 0 secs 487 ms directions/coherences Filename: 11+cbi_ep2d_bold_recon.img GroupName: Raw StimFilename: /Volumes/drobo/data/nyu/erTutorial/Etc/stimvol01.mat voxelSize=[3.0 3.0 3.0] TR=NaN framePeriod=1.2000 Dims: [64 64 22] Volumes=375 junkFrames=0 totalJunkedFrames=[0] ++++++++++++++++++++++++++ qform ++++++++++++++++++++++++++ 0.030927 -2.997722 0.112729 95.876808 -1.209577 -0.115626 -2.742910 16.015213 -2.745171 0.017175 1.209851 80.334190 0.000000 0.000000 0.000000 1.000000 qformCode: 1 ++++++++++++++++++++++++++ sform ++++++++++++++++++++++++++ -0.062857 -2.991120 0.147798 103.952255 -1.253544 -0.204434 -2.736672 22.322361 -2.725102 0.161312 1.218505 66.703751 0.000000 0.000000 0.000000 1.000000 sformCode: 1
This function also takes an output argument if you prefer to have the above information as a structure.
sInfo = scanInfo(1,'Raw');
GUI Dialogs
mrParamsDialog
usage: mrParamsDialog(paramsInfo,<titleString>,<optional arguments>)
purpose: Simple way to put up a parameter selection dialog
Optional arguments other than titleString can be given in no particular order using the string/value pair syntax or a single string containing an equal sign.
| argument | value |
|---|---|
| paramInfo | A cell array of cell arrays that contains information about parameters (see below) |
| title | An optional string containing the title you want, default is “Set Parameters” |
| buttonWidth | scale factor of the button width |
| callback | Callback function to call every time the dialog is updated |
| callbackArg | A variable to pass to the callback function |
| okCallback | A callback to call when ok is pressed |
| cancelCallback | A callback to call when cancel is pressed |
| modal | if 'modal=0' |
The first argument (paraminfo) is a cell array where each element is itself a cell array including information about a given parameter.
Each parameter must be defined by a cell array of at least 2 cells
| cell | argument | value |
|---|---|---|
| 1 | variable name | A string that will be the name of the parameter field in the output structure |
| 2 | default value | the value for this paramater, that will appear in the dialog or will be the default if mrParamsDefault is called instead of mrParamsDialog. This value could be a scalar, a string, an array or an array of strings. By default, the type of this value will define the type of uicontrol: scalars will be interpreted as numeric, arrays as array, strings as strings and cell arrays as popupmenu. |
Other cells in this array are optional arguments that can be set in no particular order. Optional arguments can be specified either as value pairs (parameter string and value) or as single strings:
| optional arguments | value |
|---|---|
| 'description' | description of the parameter that will appear in the help window. This string cannot be a single word or contain a = sign (unless it is preceded by a space). Any of the latter will be interpreted as one of the following options |
| type | Use this option to change the default uicontrol type based on the type of the default value for this parameter. Valid values are: numeric, string, array, stringarray, popupmenu, checkbox, pushbutton and statictext |
| editable | if 'editable=0', the string, scalar or array will not be editable. the default is editable=1. This option is not defined for popupmenus, checkboxes and pushbutton |
| enable | if 'enable=0', the control for this parameter will be visible, but grayed out and non-editable. the returned value will be empty for this parameter |
| visible | if 'visible=0'. the parameter will not be displayed int he dialog box. However, its default value will be output in the params structure |
| 'minmax=[a b]' | the value entered for this parameter will be restricted to the [a b] interval. a/b can be -inf/inf |
| 'incdec=[a b]' | this will append two buttons to decrease/increase the parameter value by a/b respectively |
| incdecType | 'plusMinus' draws plus and minus buttons on top of each other on the right hand side of the parameter entry field, instead of arrows on each side (the default) |
| round | if 'round=1' the user-entered parameter value is rounded to the nearest integer |
| callback | name or handle of the callback to be called. For pusbuttons, the callback must return a value that will be the value for the parameter. For other types of controls, the callback will be executed after other checks have been performed on the value and the callback cannot return an argument (these restrictions will be lift in a future version) |
| callbackArg | an argument to pass to the callback function |
| passParams | passes the current state of the params structure to the callback |
| passValue | this is not in the distributed version yet |
| passCallbackOutput | This is defaulted to 1 for pushbuttons so that the callback returns a value which sets the field (for instance if you want to set a cropRegion you can call the selectCropRegion function and that will return data that gets set. Set to 0 if your callback doesn't return any output argument (for instance if the field is just a button which brings up a visualization of some parameter) |
| contingent | name of a parameter that will control whether this parameter is enabled. The master parameter must be of the checkbox type |
| group |
Examples
You can use this function to put up a default parameter selection GUI. For example, if you just had one parameter called param which has a default value of 'paramValue', then you can do:
params=mrParamsDialog({{'param','paramValue'}});
You should make sure to add some description for the param value so that help has something in it:
params=mrParamsDialog({{'param','paramValue','This is a description of what the parameter is for'}});
If your parameter is a number and not a string, simply change the default to a numbered value
params=mrParamsDialog({{'param',0}});
If your parameter is an array or 2D matrix then simply pass that in as the default
params=mrParamsDialog({{'param',[1 2 3 4;5 6 7 8]}});
If you want to just display the values w/out allowing the user to change them set editable=0:
params=mrParamsDialog({{'param',0,'editable=0'}});
If it has a min and max value:
params=mrParamsDialog({{'param',0,'minmax=[0 10]'}});
If you want to make the value forced to be round to an integer do:
params=mrParamsDialog({{'param',0,'round=1'}});
If you want it to have arrow buttons, so that the user can increment or decrement the value easily, do:
params=mrParamsDialog({{'param',0,'incdec=[-1 1]'}});
If the variable can be one of a number of different string choice, make the default value a cell array of strings (note that the first one will be the default:
params=mrParamsDialog({{'param',{'option 1','option 2'}}});
You can also set the variable to have a button instead. In this case you will specify a callback function which will set the value of the variable. For example, if you want to have a random number set for a parameter when the user hits a button, you could do:
params=mrParamsDialog({{'randNum',0,'type=pushbutton','callback=rand','buttonString=push me to select a rand num'}});
If you want that function to receive the current parameter settings, you can do:
params=mrParamsDialog({{'randNum',0,'type=pushbutton','callback',@myFunc,'buttonString=push me','passParams=1'}});
If you want that function to receive a variable of your choosing:
params=mrParamsDialog({{'randNum',0,'type=pushbutton','callback',@myFunc,'buttonString=push me','callbackArg',myVariable}});
If you set the button to return both the params and your own variable, then your own variable will be the first argument and the second argument will be the params. Note that with the button functions you will always need to return a value.
If you have a variable that should be set as a checkbox (either 1 or 0):
params=mrParamsDialog({{'param',0,'type=checkbox'}});
If you want to have another variable contingent on this checkbox (i.e. grayed out if the checbox is zero), then do:
params=mrParamsDialog({{'param',0,'type=checkbox'},{'otherVar','value','contingent=param'}});
If you have a set of parameters that depends on another, you can do:
paramInfo = {...
{'param',1,'round=1','incdec=[-1 1]','minmax=[0 2]'},...
{'otherVar',{{'Set1 value1','Set1 value2'},{'Set2 Value1'}},'contingent=param'}};
params=mrParamsDialog(paramInfo);
You can also make a group of parameters that works similar to parameters that are contingent on each other. For example if you want to set differnet parameters for each of two scans and return an array of settings, you could do:
paramInfo = {...
{'scanNum',1,'round=1','incdec=[-1 1]','minmax=[1 2]'},...
{'groupedNumeric',{1 10},'group=scanNum','type=numeric','incdec=[-1 1]'},...
{'groupedString',{'firstScan' 'secondScan'},'group=scanNum','type=String'},...
{'groupedCheckbox',{0 1},'group=scanNum','type=checkbox'}};
params=mrParamsDialog(paramInfo);
For more than one parameter, add more things into the array. Here is a full example:
paramInfo = {...
{'Title','Default Title','This is the title'},...
{'check',1,'type=checkbox','This is a checkbox to do something or not'},...
{'groupName',{'Raw','MotionComp'},'The group to select from'},...
{'scanNum',1,'minmax=[1 10]','incdec=[-1 1]','The scan number to use'},...
}
params = mrParamsDialog(paramInfo);
mrParamsDialog will return with a params structure if the user hit ok, and [] if the user hit cancel. The field params.paramInfo will be set to the passed in paramInfo.
You can also use mrParamsDialog to make a modeless dialog (i.e. one which stays up, doesn't have ok/cancel buttons and every time a parameter is changed calls your callback function). This is usefull for using mrParamsDialog as an interface control panel:
mrParamsDialog(paramsInfo,'Controls',[],@myCallback,optionalArgument);
The function myCallback should be defined to accept as the first argument the params structure and as the second argument the optionalArgument (if any–if you don't specify an optionalArgument it won't be passed).
mrParamsDefault
usage: params = mrParamsDefault(paramsInfo);
purpose: Gets default parameter settings from the paramsInfo structure used by mrParamsDialog
| argument | value |
|---|---|
| paramInfo | A cell array of cell arrays that contains information about parameters |
This function takes the same paramsInfo structure as mrParamsDialog, but instead of putting up a dialog simply sets all the defaults and returns the params structure. This is useful for writing functions such that they accept the “defaultParams=1' flag.
params = mrParamsDefault(paramsInfo);
mrParamsSet
usage: mrParamsSet(params)
purpose: Used when running mrParamsDialog with a callback. Pass in modified params and this function will change the mrParamsDialog GUI to reflect those changes.
| argument | value |
|---|---|
| params | The parameter structure returned to the callback |
mrParamsDialogSelectScans
usage: paramsInfo = mrParamsDialogSelectScans(v,groupNum,<paramsInfo>,<includeList>)
purpose: create paramsInfo structure for use with mrParamsDialog that allows selection of scans.
| argument | value |
|---|---|
| v | view variable |
| groupNum | Number of group to select scans from |
| paramsInfo | An already created paramsInfo structure to append these parameters to |
| includeList | List of scans to default to include |
To put up a selection for scans, you can do:
v = newView; paramsInfo = mrParamsDialogSelectScans(v,1); params = mrParamsDialog(paramsInfo); if ~isempty(params) scanNums = find(params.include); end
If you want to only have one scan selected (say scan 3) at start up:
paramsInfo = mrParamsDialogSelectScans(v,1,{},3);
Nifti Headers
mrUpdateNiftiHdr
This function is no longer necessary. It updates the nifti headers in the mrSession from the nifti headers in the files. Should be run if you already have run mrInitRet and then make a change to the nifti headers using mrAlign. But, usually you will use the export to mrLoadRet-4.5 feature of mrAlign so you don't need to use this function.
mrFixNiftiHdr
usage: mrFixNiftiHdr(<forceUpdate>,<copyFrom>)
purpose: Helper function to fix problems with nifti qform/sforms
| argument | value |
|---|---|
| forceUpdate | Set to 1 to do update without asking |
| copyFrom | group/scan from which to copy the nifti header |
Looks for inconsistencies between the dimensions of the voxels listed in the header and that calculated in the qform. If there is an inconsistency it will give you the option to copy the header from the raw header to fix it. Often these problems occur after using FSL to do motion correction since it doesn't preserve the nifti header properly. The symptom is that your voxel sizes get set to strange numbers when you use cbiWriteNifti since that code calculates voxel dimensions from the qform matrix when you save.
You can use forceUpdate to force mrFixNiftiHdr to copy the headers for a set of scans. If forceUpdate is a single number, it means you want to update every scan in a group:
mrFixNiftiHdr(2)
Or it can be a list of scans/group pairs, for instance i f you want to fix scan 1 and 4 from group 3 and 2 respectively:
mrFixNiftiHdr([1 3;4 2]);
If you want to specify the nifit header that should be used to copy from to fix the headers, you can do (e.g. if you wanted to copyFrom the header from scan 3, group 1):
mrFixNiftiHdr([1 3;4 2],[3 1]);
mrSetNiftiTransforms
usage: mrSetNiftiTransforms(scanAndGroup,qform44,sform44)
purpose: Function to allow you to set the qform and sform of scans. Useful for fudging things when the nifti files are broken.
| argument | value |
|---|---|
| scanAndGroup | An array that specifies the scan and groups to be updated |
| qform44 | The qform to set the transform to |
| sform44 | The sform to set the transform to |
If you want to update scans 1 and 4 from group 3 and 2.
mrSetNiftiTransforms([1 3;4 2],eye(4),sform);
To leave one transform unchanged do
mrSetNiftiTransforms([1 3;4 2],eye(4),[]);
Viewers
There are a few functions that can serve to view data or surfaces that are usually called through the menu items, but can be used directly by themselves.
mlrDisplayEPI
usage: mlrDisplayEPI(<v/data/filename>,<groupName>)
purpose: Display EPIs for the specified group. Same a plot menu item..
| argument | value |
|---|---|
| v/data/filename | view variable or data structure of filename |
| groupName | Name of group to display |
Can be called with a view:
v = newView; mlrDisplayEPI(v,'MotionComp')
Or with a volume
data = cbiReadNifti('epiImages.hdr');
mlrDisplayEPI(data);
Or with a filename
mlrDisplayEPI('epiImages.hdr');
mrFlatViewer
usage: params = mrFlatViewer(flat,<outer>,<inner>,<curv>,<anat>,<viewNum>)
purpose: Displays a flattened patch.
| argument | value |
|---|---|
| flat | Name of flat file to display |
| outer | Name of outer surface |
| inner | Name of inner surface |
| curv | Name of curvature file |
| anat | Name of 3D anatomy file |
| viewNum | View number |
Flat can be a name of a file:
mrFlatViewer('jg_left_MT_flat');
Or can be a structure that specifies the location of a patch defined by a point/radius:
flat.path = pwd; flat.parentSurfaceName = 'jg_left_WM'; flat.startPoint = [200 50 100]; flat.radius = 50; params = mrFlatViewer(flat);
This function is used both to display flat files and to get user defined parameters.
mrSurfViewer
usage: mrSurfViewer(outerSurface,<outerCoords>,<innerSurface>,<innerCoords>,<curv>,<anat>)
purpose: Displays a surface.
| argument | value |
|---|---|
| outerSurface | Name of outer display surface |
| outerCoords | Name of outer coordinate surface |
| innerSurface | Name of inner display surface |
| innerCoords | Name of inner coordinate surface |
| curv | Name of curvature file |
| anat | Name of 3D anatomy file |
Additional functions (might change)
The following functions are functions that might change