Table of Contents
Overview
If you do not have orientation information for your MRI data (for example if you do not have your data in Nifti format) you may still use MLR. But, you will need to manually set the orientation information - here are some steps for how to do this. This assumes that you have a “canonical” 3D anatomy taken in a separate session. In your functional session, along with your epi images you will need a 2D inplane image taken with exactly the same orientation as the epi images. Note that it can have a different resolution (e.g. the inplanes can be 1.5×1.5×3 for epi's that are 3x3x3). Assuming you have these images then you will do the following:
- Set the 3D volume qform
- Set the 2D inplane qform
- Roughly align the 2D to the 3D
- Set the epi qform
After you are done with these steps (detailed below), you will be able to do the normal MLR processing steps, including using mrAlign to precisely align your 2D inplane to the 3D volume.
Set the 3D volume qform
Load your 3D anatomy into mlrVol. Assuming that the variable d holds your 3D data:
mlrVol(d)
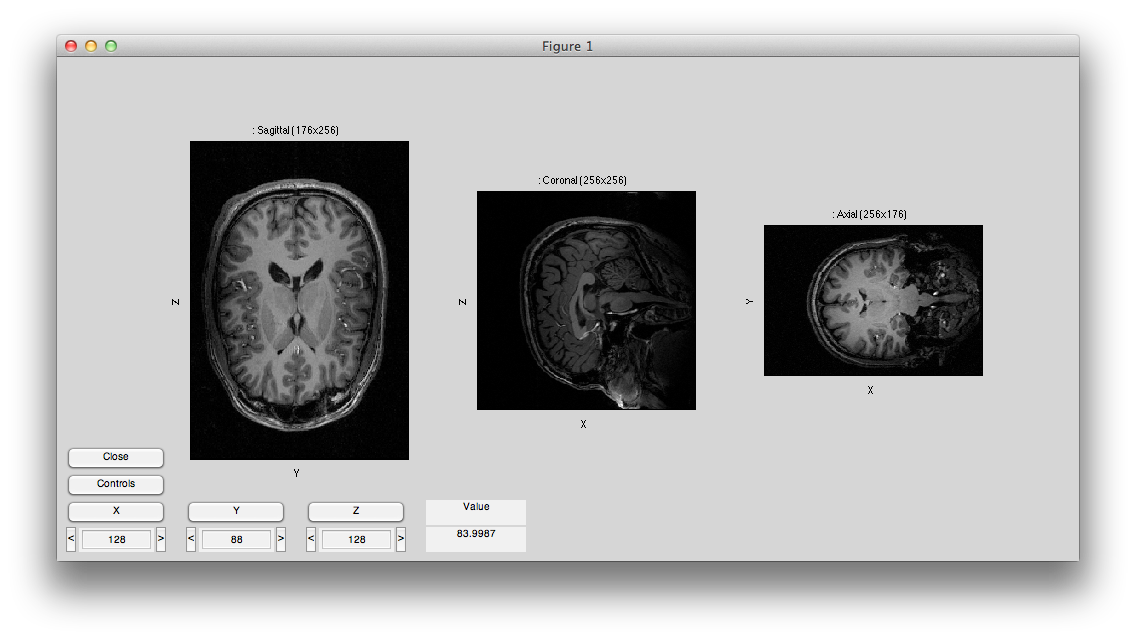
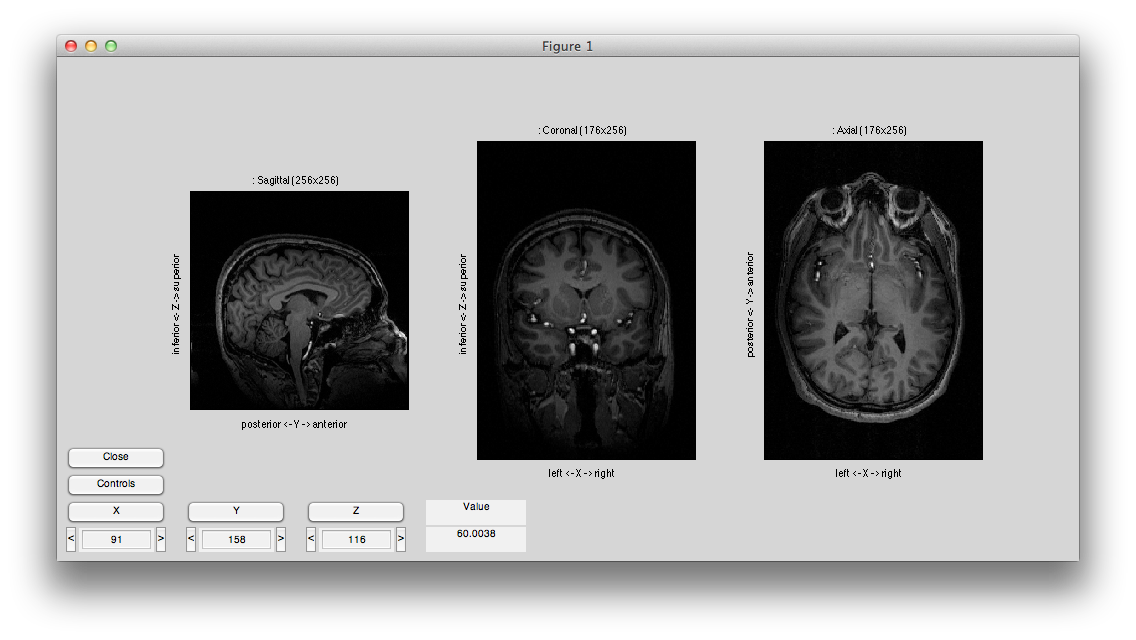
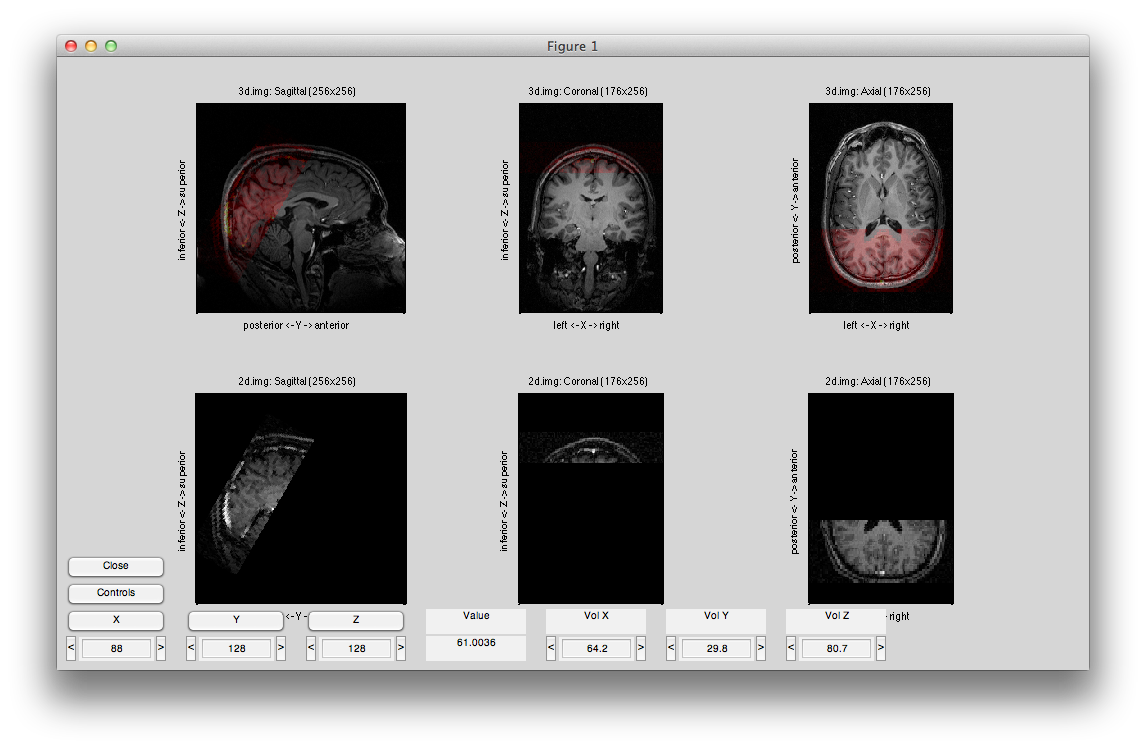
You should see something like this:
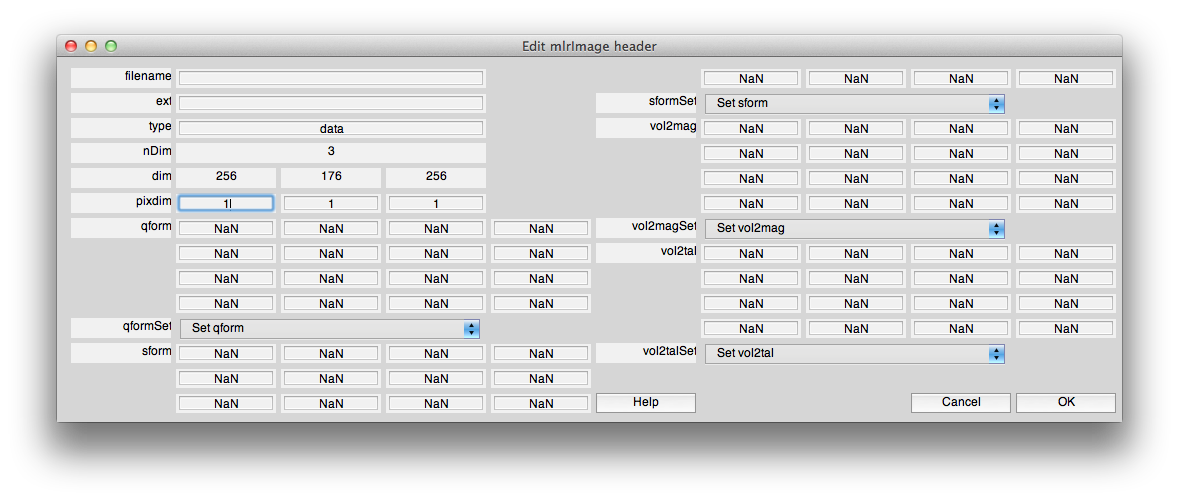
First set the pixdim for the volume. Hit the controls button and then edit header, and input the voxel dimensions in the pixdim field. This volume is 1x1x1 so we enter 1 1 1:
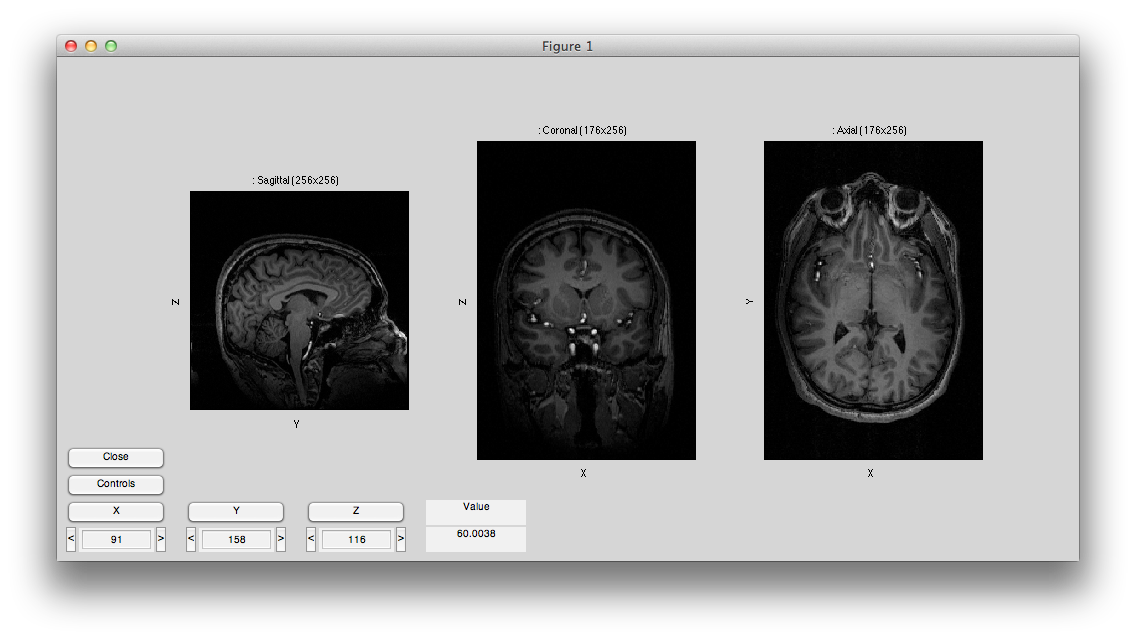
Now hit controls again and then try different swaps and flips until your volume is in the canonical orientation, with the leftmost subplot showing the sagittal image with the nose to the right, the center subplot showing the coronal image with left on the left and the rightmost subplot showing an axial image with left on the left and the eyes at the top. It should look like this:
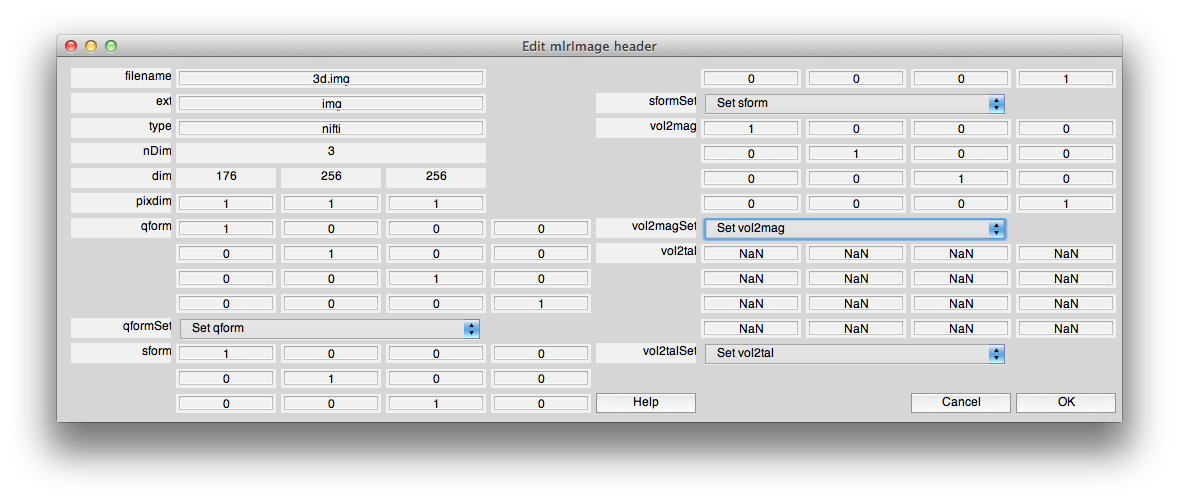
Now, click edit header and use the “Set qform” control to set the qform to “identity”. Usually we set the sform (and the vol2mag) to the qform for canonical volumes (by clicking set base coordinate frame in mrAlign). You may want to do that now, by setting the sform and vol2mag to identity. The header should look like the following:
Labels (e.g. left/right) should appear on all of the axis, like the following. Make sure that they are all correct:
If they are all correct, then hit Controls and then “Save volume”. Save the volume. It now has a valid nifti header with qform information.
Set the 2D inplane qform
Now you will need to set the qform (orientation in the magnet for the 2D inplane. You do this by setting the qform to identity like for the 3D volume. Then you will roughly align by hand the 2D inplane to the 3 Load your session 2D anatomy. This should be an anatomy that is in the same orientation with the same number of slices as your EPI, but does not have to have the same inplane resolution (i.e. usually it has a higher resolution). Follow the same procedure as above to set its xform.
- Load the 2D anat into mlrVol
- Set its pixdim
- Swap and flip to put it into a canonical orientation
- Set the qform to identity
- You may want to check the pixdim again to make sure it is correct, especially if you are not using isotropic voxels (make sure that the correct dimensions have the right size).
- Save
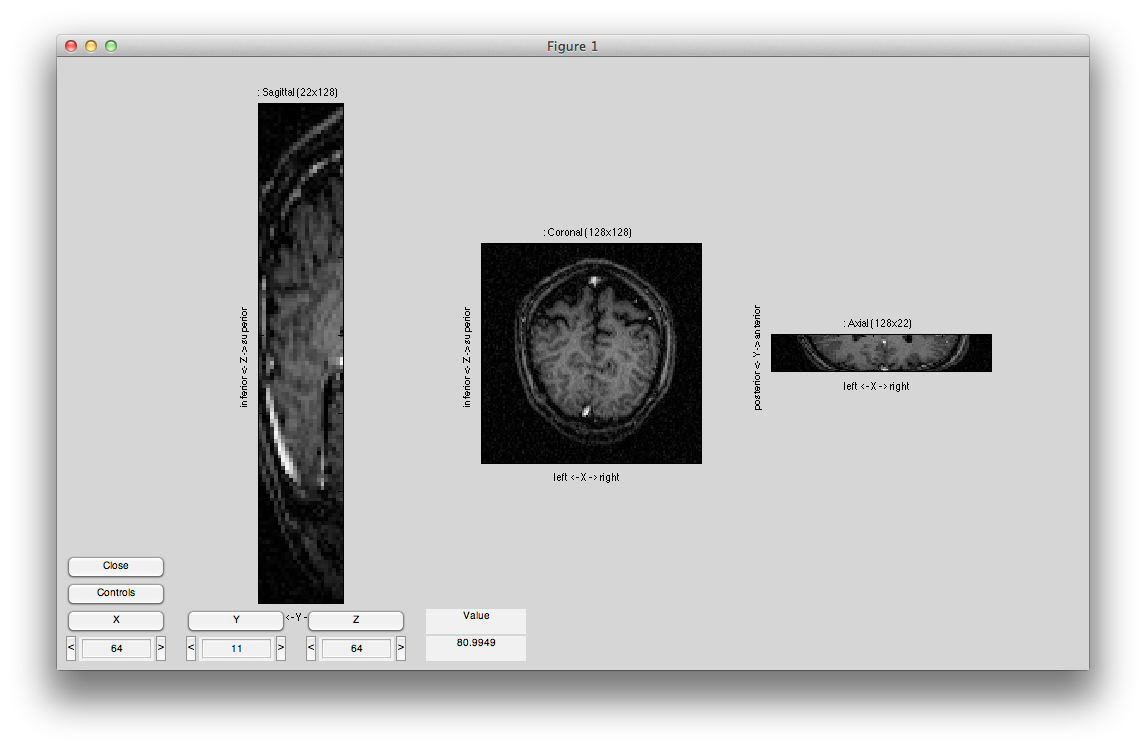
After you are done, it should look something like this:
Rough manual alignment of 2D to 3D
Now, you should do a rough alignment of the 2D to the 3D. You will use mrAlign afterwords to do a final alignment, so this does not have to be very accurate - it just has to have the slices in roughly the right orientation and location so that mrAlign has a good starting place. To do the rough alignment, load the 3D and the 2D that you have created in the above steps into mlrVol
mlrVol('3d.nii','2d.nii');
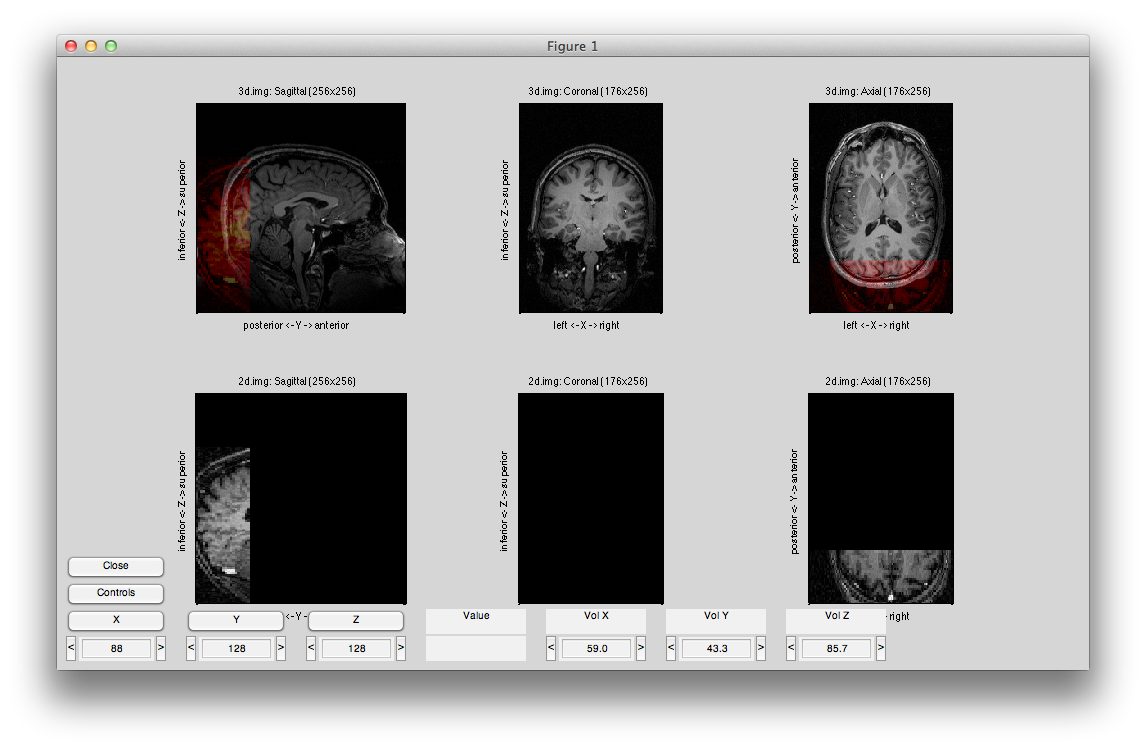
The alignment, will probably look something like this:
You may want to align the centers of the two images to start with. To do so, click Controls and then “Set to Identity”. It will look like the following:
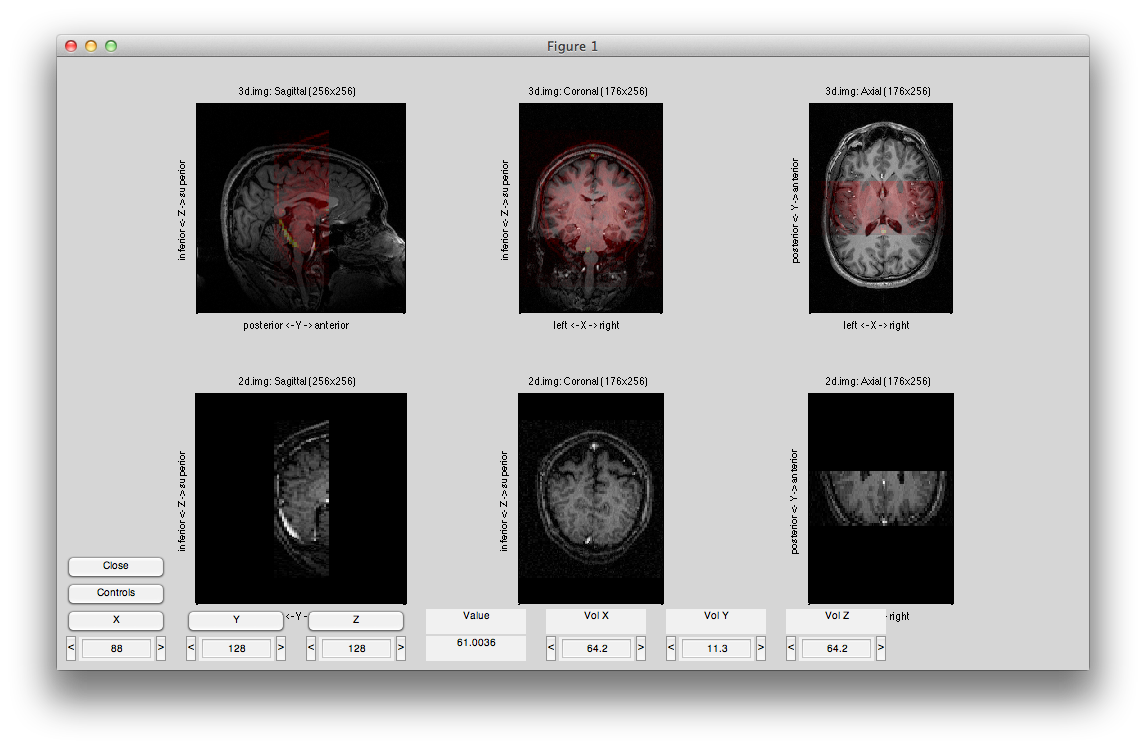
Now use rotations and shifts to roughly align the 2D to the 3D. When you are done, it should look like something like the following:
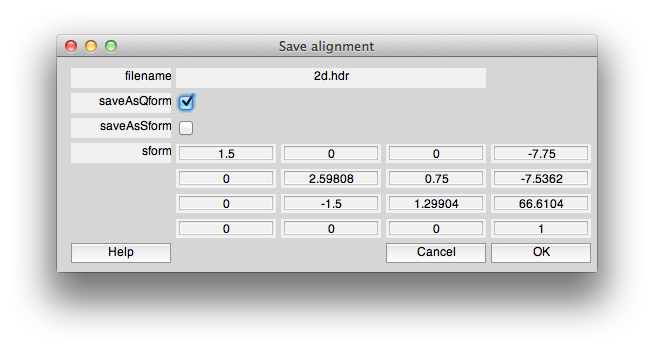
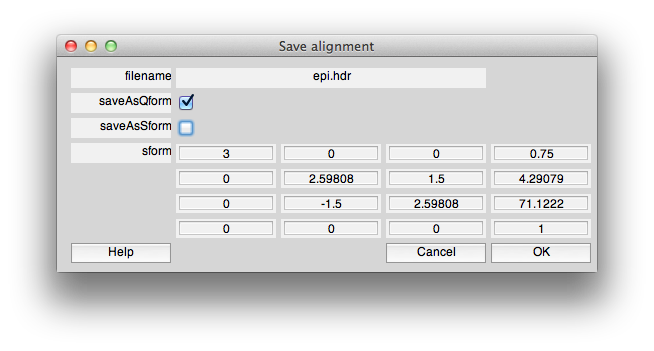
When you are done, you will save the alignment to the qform. To do this click “Save alignment” and then choose the filename and when you see the following dialog, make sure that saveAsQform is checked and saveAsSform is unchecked:
Set the epi qform
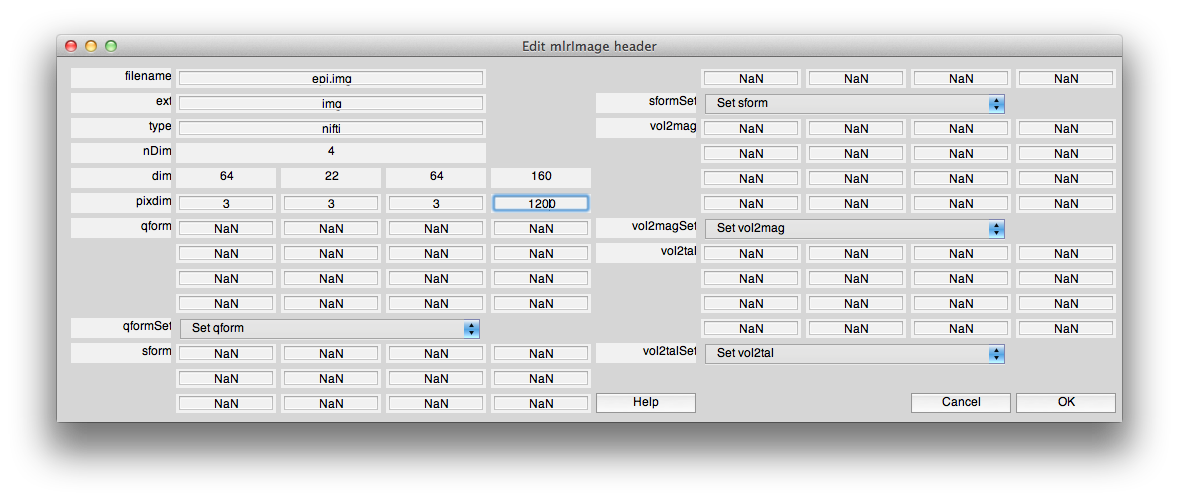
Now, load your epi image data in mlrVol and set the pixdim. Note that the fourth element in the pixdim should be the frame period (volume TR) in ms. It should look something like the following:
Save the volume. Now we will check the alignment with the 2D anatomy. This will automatically get calculated to assume that the volumes have the same center:
mlrVol 2d.nii epi.nii
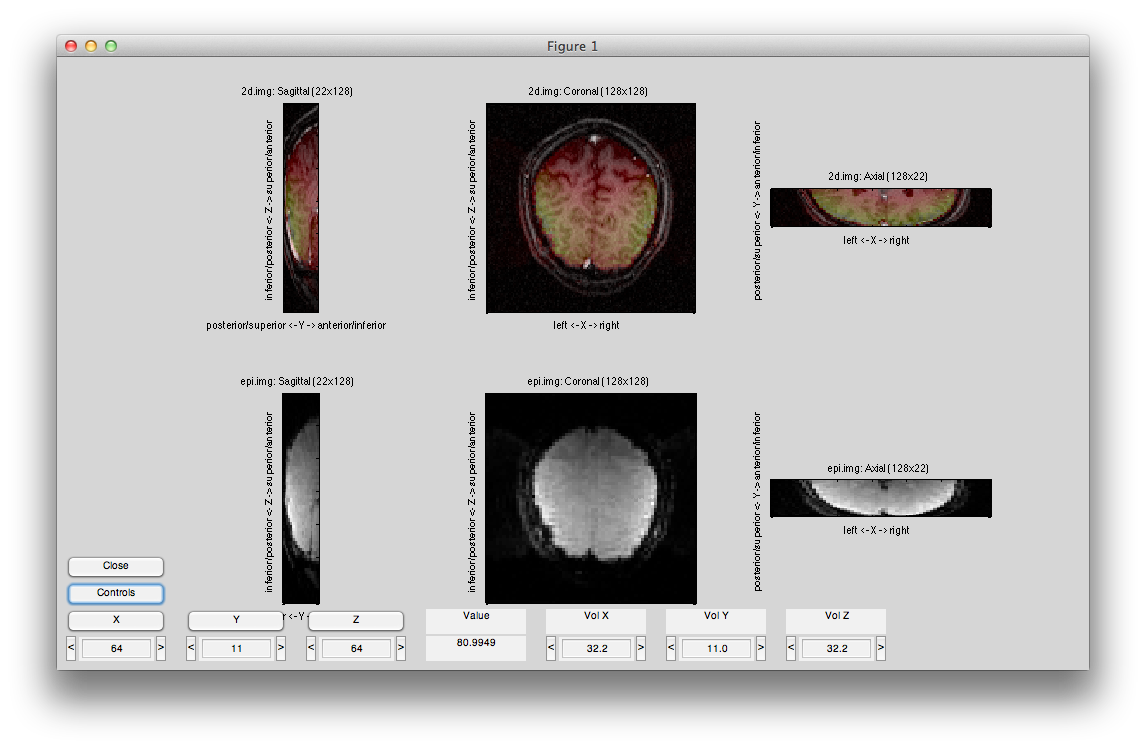
It should look like the following:
If you check the vol2vol in the controls, it should have a straight-forward form. For example, if your 2d has twice the inplane resolution as the epi image then you will see a vol2vol like:
2 0 0 0.5 0 2 0 0.5 0 0 1 0 0 0 0 1
Note the 2's on the diagonal represent the doubling of the inplane resolution, and the shift of 0.5 is to align the centers of the first voxels.
You can now save your alignment to the qform.